Why someone with a healthy lifestyle suffers from a heart attack? Or alternatively, why someone who smokes, does not exercise and have a poor diet remains completely healthy? The answer to these critical questions lies at the interface between genetics and the environment. Our lab addresses them using modern human genetic approaches and high-throughput genomic technologies.
Using genetics and functional genomics to enable precision medicine and drug development in cardiovascular and blood diseases
Opportunities
Opportunities exist for talented and motivated individuals (students, postdocs, fellows, programmers, bioinformaticians, and (bio)statisticians) to join our research activities. Please send email inquiries to Guillaume.Lettre [at] mhi-humangenetics.org .
Equity, Diversity and Inclusion
We value equity, diversity and inclusion because the goal of the lab is to improve health for all individuals.
News
- 2022-01-13. Prof. Guillaume Lettre is awarded a tier 1 Canada Research Chair in Heart and Blood Diseases Genetics.

- 2021-09-15. The lab is an awardee of the Impact of Genomic Variation on Function (IGVF) Consortium.

- 2020-09-04. Dr Guillaume Lettre talks about the genetics of blood.

- 2019-09-10. Dr Guillaume Lettre is elected to the Royal Society of Canada College of New Scholars, Artists and Scientists.

- 2019-06-11. Dr Guillaume Lettre talks about polygenic risk score relevance, which was applied to predict likelihood of heart disease in French Canadian.

- 2018-10-11. Dr Guillaume Lettre is the winner of the André-Dupont 2018 Award.

- 2017-02-02. Dr Guillaume Lettre and Ken Sin Lo‘s work on a large-scale international study about the genetics of adult height is now published in Nature.


- 2016-06-29. Publication of three ExomeChip articles in the AJHG by the Blood-Cell Consortium led by Dr Guillaume Lettre. Congratulations to PhD student Nathalie Chami, lead co-author on 2 of the 3 papers.
- 2015-05-01. Congratulations to Nathalie Chami and Samuel Lessard who were awarded PhD scholarships from FRQS and CIHR! Geneviève Galarneau, former PhD student in the Lettre lab, also received a CIHR postdoctoral fellowship to continue her training at Mount Sinai (New York).
- 2014-04-28. Dr Guillaume Lettre identifies genetic mutations involved in human blood diseases.

- 2013-10-11. An article with Dr Guillaume Lettre and Samuel Lessard as co-authors made the cover of Science.

- 2013-10-04. Lip-sync!

- 2013-07-18. SCD findings highlighted as one of the hottest studies in Blood.

- 2012-12-12. Dr Guillaume Lettre receives a Doris Duke Innovations in Clinical Research Award.

- 2012-01-23. Dr Guillaume Lettre talks about height.

- 2010-11-24. Dr Guillaume Lettre receives a Canada Research Chair.

- 2010-09-30. Dr Guillaume Lettre‘s laboratory publishes an important study in Nature on the genetics of adult height.



Projects
Endothelial cell genomics
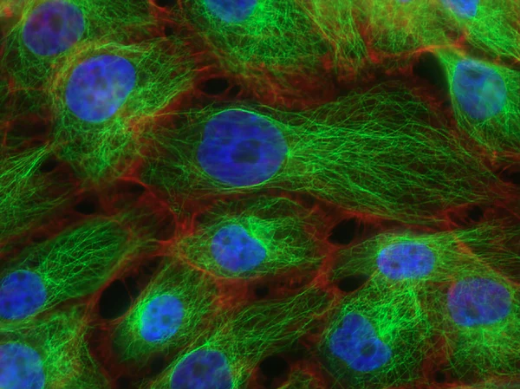
Endothelial cells form the inner layer of blood vessels and play a critical role to maintain cardiovascular health. They form a selective barrier, respond to inflammation, and control the vascular tone and hemostatic factors.
We have developed a comprehensive program to characterize endothelial cells’ responses to different stimuli, including transcriptomic (RNA-sequencing), epigenomic (open chromatin regions, ATAC-sequencing) and chromosome conformation (Hi-C) changes.
We are also using CRISPR/Cas9 genome editing screens to interrogate how genes and non-coding regulatory elements control endothelial cell functions. Current projects focus on atherosclerosis and hypertension.
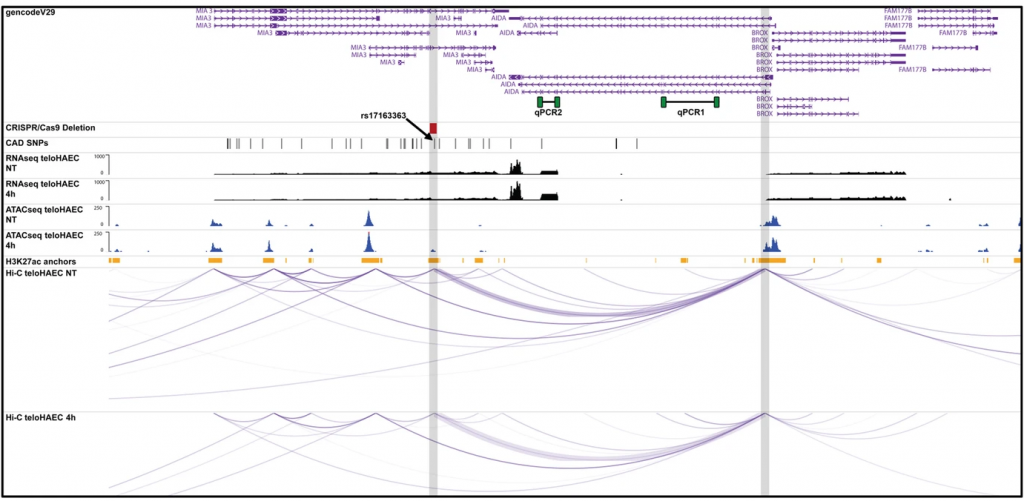
Figure 5a. Lalonde et al., Genome Biology, 2019
Graphical representation of the transcriptomic, epigenomic, and 3D conformation data at the AIDA coronary artery disease (CAD)-associated locus. The CRISPR/Cas9 deletion is indicated in red and both qPCR assays are represented in green. We added gray vertical bars to highlight the TNFα-sensitive open chromatin peak (left) and the AIDA promoter (right). To improve visualization, we also increased the width of the arcs linking both elements (purple).
Cardiovascular disease genetics
Cardiovascular diseases (CVDs) are the main cause of death in the Western world. In Canada, CVDs affect 1.3 million individuals and cost nearly $22 billions (direct and indirect costs) to the Canadian economy.
Beyond the traditional risk factors (e.g. lipids, hypertension, obesity), familial history (genetics) is also predictive of an individual’s risk to develop CVDs. We used comprehensive genetic datasets (including whole-genome sequencing) in very large sample sizes (>700,000 participants) to discover genetic risk factors for CVDs.
These discoveries guide our functional experiments, and also allow us to optimize tools to predict CVDs risk. The current focus in the lab is on atherosclerosis and atrial fibrillation.
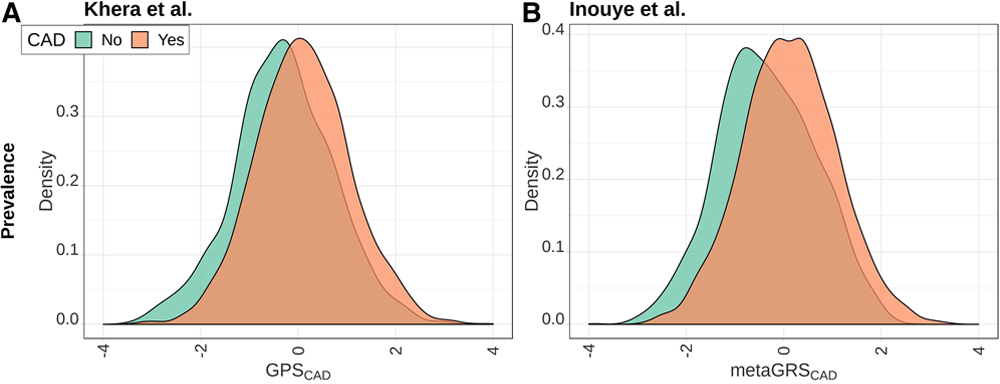
Figure 2. Wünnemann et al., Circulation: Genomic and Precision Medicine, 2019
Distributions of GPSCAD and metaGRSCAD in the Montreal Heart Institute (MHI) Biobank phase 2 cohort. Distributions of the normalized polygenic risk score from Khera et al11 (GPSCAD, left column) and Inouye et al12 (metaGRSCAD, right column) in the MHI Biobank phase 2 data for prevalent (A and B), incident (C and D), and recurrent (E and F) coronary artery disease (CAD) events.
Blood-cell genetics
Blood is mostly composed of plasma and blood cells and plays a major role in a variety of functions involved in general human health: it transports oxygen, nutrients and hormones to tissues, removes waste, performs immunological functions and contributes tissue damage repair through coagulation.
The main three blood cell types carry out most of these activities: red blood cells transport oxygen, white blood cells coordinate some of the immune responses, and platelets are the bricks that form blood clots to prevent excessive bleeding.
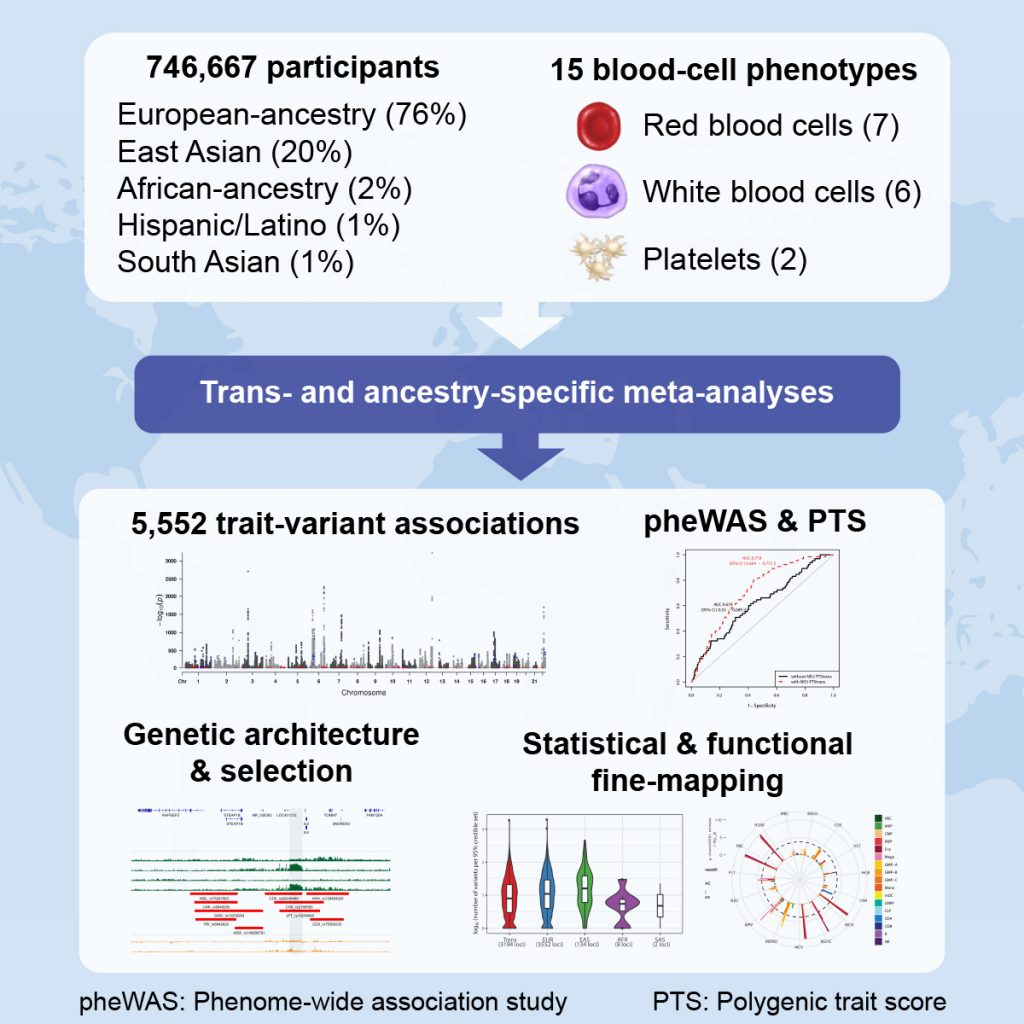
All of these cell types originate through proliferation and differentiation from common precursors (hematopoietic stem cells). An aberrant number, size or feature of the three main blood cell types characterizes multiple human diseases. We created the international Blood-Cell Consortium to study how genetic factors control hematopoiesis.
We combine our genetic analyses with genomic (e.g. expression quantitative trait loci (eQTL)) and genome editing (e.g. CRISPR/Cas9) approaches to identify genes that regulate blood-cell phenotypes. Furthermore, we are also interested in exploring whether blood-cell phenotypes can be used as predictive biomarkers of hematological and cardiovascular diseases.
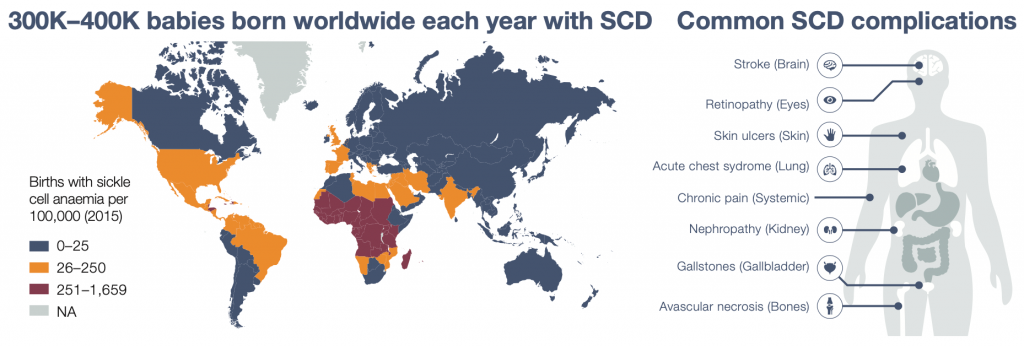
Sickle cell disease
Sickle cell disease (SCD) is one of the most common monogenic diseases in the world. It is particularly prevalent in countries where malaria is endemic, as carrying one copy of the SCD mutation confers protection from malaria infection.
SCD presents an extremely heterogenous clinical course, with some patients being only mildly affected whereas others suffer devastating complications (e.g. stroke, renal failure). To understand this heterogeneity, we have been focusing on the genetic regulation of fetal hemoglobin levels, a strong modifier of SCD severity.
We are now moving towards other phenotypes and measures of severity to further understand the pathophysiology of SCD. We are also interested in developing new therapies for SCD.